If there is one lesson to learn from comparative genomics, it is to never underestimate the bacteria. Bacteria are verily the engines of protein diversity and have provided some of the most remarkable insights on the origins of various pathways and systems. In our next release on the ubiquitin-mediated signaling/ tagging/ protein turnover pathway, we address the origins and roles of several treble clef domains, including the RING finger, found in the eukaryotic ubiquitin pathway. This study also answers some tricky questions raised in a previous post.
Q1. Do any species have the entire complement of both the ubiquitin and pupylation based protein turnover/tagging system?
Indeed, some actinobacteria such as Frankia and planctomycetes such as Pirellula staleyi contain both the entire complement of the ubiquitin system and pupylation. The list of species with both systems is only expected to grow as more such genomes are sequenced. The discovery of the Frankia RING finger was a bit complicated. It turns out that the RING finger is encoded in the opposite strand to the sequence that is submitted to the database and whats more the RING is sandwiched between an E1 and E2. For details, peruse our supplement.
Q2. If so, how is the labor of protein turnover divided between pupylation and ubiquitination?
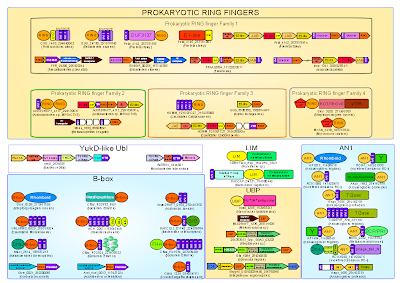
This far we can only make a reasonable guess, but this is where experiments will reveal more. There is, however, a curious aspect to the prokaryotic RING-finger containing Ub-systems. Many of the core Ub-pathway proteins have transmembrane (TM) helices. In Frankia, both the JAB deubiquitinase and the RING are fused to transmembrane helices. In Pirellula, the RING finger while containing a TM helix is also fused to a distinct domain prototyped by DUF3137, a potential solute sensor. This suggests that a major fraction of the prokaryotic RING domains might have functions related to either regulation or modification of membrane-associated proteins. While the pupylation system of actinobacteria is cytoplasmic, the Pirellula Pup ligase, like other planctomycete versions, has 4 transmembrane helices inserted within the core domain, suggesting that in some species, pupylation has a strong membrane component. Study the architectures and associations of these Ub-systems here
Q3. In light of the discovery in Caldiarchaeum, what can we say about the origins of the eukaryotic Ub-system?
The Caldiarchaeum Ub-system is remarkable in that each of its Ub-system components is very closely related in sequence and structure to the corresponding eukaryotic version. For example, the E1 protein of this archaeon contains a C-terminal Ub-fold UFD domain (that was only detected in eukaryotes to date). However, this far no other archaeon contains a complete complement of the Ub-system. Further bacterial versions of the complete Ub-system are also sporadically distributed, suggesting a strong component of lateral transfer in the dissemination of this system across prokaryotes. Therefore, we cannot be certain if the eukaryotic Ub-system emerged from a Caldiarchaeum-like system in the archaeal symbiont during eukaryogenesis. Indeed, such systems might be present in as yet un-sampled bacteria suggesting that it is not unlikely that eukaryotes acquired such a system from the primary bacterial symbiont or even via an independent lateral transfer of the operon from yet another prokaryote.
Based on this and our previous study on the ubiquitin system, we can now confidently state that systems resembling eukaryotic Ub-conjugation systems were put together to different degrees in prokaryotes during the diversification of various biosynthetic and regulatory pathways. With regards to the proteasomal association, while the core proteasomal apparatus is of archaeal origin, it is also present in various bacteria of which some possess the complete core Ub-system. Hence, it is possible that this connection between the Ub system and the proteasome developed either in bacteria or archaea, and was merely retained in eukaryotes which vertically inherited their core proteasomal complex from the archaea.
Wait.. there much more to this story. Feel free to access the paper here and browse through the extensive supplement.